The regulatory T cells (Tregs), formerly known as suppressor T cells, are a subpopulation of T cells that modulate the immune system, maintain tolerance to self-antigens, and prevent autoimmune disease. Tregs are immunosuppressive and generally suppress or downregulate induction and proliferation of effector T cells.[1] Tregs express the biomarkers CD4, FOXP3, and CD25 and are thought to be derived from the same lineage as naïve CD4 cells.[2] Because effector T cells also express CD4 and CD25, Tregs are very difficult to effectively discern from effector CD4+, making them difficult to study. Recent research has found that the cytokine TGFβ is essential for Tregs to differentiate from naïve CD4+ cells and is important in maintaining Treg homeostasis.[3]
Mouse models have suggested that modulation of Tregs can treat autoimmune disease and cancer and can facilitate organ transplantation[4] and wound healing.[5]
Their implications for cancer are complicated. Tregs tend to be upregulated in individuals with cancer, which suggest that cancer is autoimmune disease. They seem to be recruited to the site of many tumors. Studies in both humans and animal models have implicated that high numbers of Tregs in the tumor microenvironment is indicative of a poor prognosis. Furthermore, Tregs are thought to suppress tumor immunity, thus hindering the body’s innate ability to control the growth of cancerous cells.[6] Recent immunotherapy research in conventional medicine is studying how regulation of T cells could possibly be utilized in the treatment of cancer.[7]. In holistic oncology, we already know important Treg cells for cancer reversal.
Generalities
T regulatory cells are a component of the immune system that suppress immune responses of other cells. This is an important “self-check” built into the immune system to prevent excessive reactions. Regulatory T cells come in many forms with the most well-understood being those that express CD4, CD25, and FOXP3 (CD4+CD25+ regulatory T cells). These “Tregs” are different from helper T cells.[8] Another regulatory T cell subset is Treg17 cells.[9] Regulatory T cells are involved in shutting down immune responses after they have successfully eliminated invading organisms, and also in preventing autoimmunity.[10]
CD4+ Foxp3+ CD25(high) regulatory T cells have been called “naturally occurring” regulatory T cells[11] to distinguish them from “suppressor” T cell populations that are generated in vitro. Additional regulatory T cell populations include Tr1, Th3, CD8+CD28-, and Qa-1 restricted T cells. The contribution of these populations to self-tolerance and immune homeostasis is less well defined. Foxp3 can be used as a good marker for mouse CD4+CD25+ T cells, although recent studies have also shown evidence for Foxp3 expression in CD4+CD25- T cells. In humans, Foxp3 is also expressed by recently activated conventional T-cells and thus does not specifically identify human Tregs.[12]
Source: Progenitor cells
All T cells derive from progenitor cells in the bone marrow, which become committed to their lineage in the thymus. All T cells begin as CD4-CD8-TCR- cells at the DN (double-negative) stage, where an individual cell will rearrange its T cell receptor genes to form a unique, functional molecule, which they, in turn, test against cells in the thymic cortex for a minimal level of interaction with self-MHC. If they receive these signals, they proliferate and express both CD4 and CD8, becoming double-positive cells. The selection of Tregs occurs on radio-resistant hematopoietically-derived MHC class II-expressing cells in the medulla or Hassal’s corpuscles in the thymus. At the DP (double-positive) stage, they are selected by their interaction with the cells within the thymus, begin the transcription of Foxp3, and become Treg cells, although they may not begin to express Foxp3 until the single-positive stage, at which point they are functional Tregs. Tregs do not have the limited TCR expression of NKT or γδ T cells; Tregs have a larger TCR diversity than effector T cells, biased towards self-peptides.
The process of Treg selection is determined by the affinity of interaction with the self-peptide MHC complex. Selection to become a Treg is a “Goldilocks” process; a T cell that receives very strong signals will undergo apoptotic death; a cell that receives a weak signal will survive and be selected to become an effector cell. If a T cell receives an intermediate signal, then it will become a regulatory cell. Due to the stochastic nature of the process of T cell activation, all T cell populations with a given TCR will end up with a mixture of Teff and Treg – the relative proportions determined by the affinities of the T cell for the self-peptide-MHC. Even in mouse models with TCR-transgenic cells selected on specific-antigen-secreting stroma, deletion or conversion is not complete.
Foxp3+ Treg generation in the thymus is delayed by several days compared to Teff cells and does not reach adult levels in either the thymus or periphery until around three weeks post-partum. Treg cells require CD28 co-stimulation and B7.2 expression is largely restricted to the medulla, the development of which seems to parallel the development of Foxp3+ cells. It has been suggested that the two are linked, but no definitive link between the processes has yet been shown. TGF-β is not required for Treg functionality, in the thymus, as thymic Tregs from TGF-β insensitive TGFβRII-DN mice are functional.
Self and non-Self
The immune system must be able to discriminate between self and non-self. When self/non-self discrimination fails, the immune system destroys cells and tissues of the body and as a result causes autoimmune diseases. Regulatory T cells actively suppress activation of the immune system and prevent pathological self-reactivity, i.e. autoimmune disease. The critical role regulatory T cells play within the immune system is evidenced by the severe autoimmune syndrome that results from a genetic deficiency in regulatory T cells (IPEX syndrome – see also below).
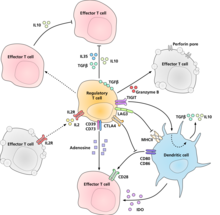
Diagram of regulatory T cell, effector T cells and dendritic cell showing putative mechanisms of suppression by regulatory T cells.
The molecular mechanism by which regulatory T cells exert their suppressor/regulatory activity has not been definitively characterized and is the subject of intense research. In vitro experiments have given mixed results regarding the requirement of cell-to-cell contact with the cell being suppressed. The following represent some of the proposed mechanisms of immune suppression:
Regulatory T cells produce a number of inhibitory cytokines. These include Transforming growth factor beta,[13] Interleukin 35,[14] and Interleukin 10.[15] It also appears that regulatory T cells can induce other cell types to express interleukin-10.[16] Regulatory T cells can produce Granzyme B, which in turn can induce apoptosis of effector cells. Regulatory T cells from Granzyme B deficient mice are reported to be less effective suppressors of the activation of effector T cells.[17] Reverse signalling through direct interaction with dendritic cells and the induction of immunsuppressive indoleamine 2,3-dioxygenase.[18] Signalling through the ectoenzymes CD39 and CD73 with the production of immunosuppressive adenosine.[19][20] Through direct interactions with dendritic cells by LAG3 and by TIGIT.[21][22] Another control mechanism is through the IL-2 feedback loop. Antigen-activated T cells produce IL-2 which then acts on IL-2 receptors on regulatory T cells alerting them to the fact that high T cell activity is occurring in the region, and they mount a suppressory response against them. This is a negative feedback loop to ensure that overreaction is not occurring. If an actual infection is present other inflammatory factors downregulate the suppression. Disruption of the loop leads to hyperreactivity, regulation can modify the strength of the immune response.[23] A related suggestion with regard to interleukin 2 is that activated regulatory T cells take up interleukin 2 so avidly that they deprive effector T cells of sufficient to avoid apoptosis.[24] A major mechanism of suppression by regulatory T cells is through the prevention of co-stimulation through CD28 on effector T cells by the action of the molecule CTLA-4.[25]
Induced regulatory T cells (iTreg)
Induced regulatory T (iTreg) cells (CD4+ CD25+ Foxp3+) are suppressive cells involved in tolerance. iTreg cells have been shown to suppress T cell proliferation and experimental autoimmune diseases. These cells include Treg17 cells. iTreg cells develop from mature CD4+ conventional T cells outside of the thymus: a defining distinction between natural regulatory T (nTreg) cells and iTreg cells. Though iTreg and nTreg cells share a similar function iTreg cells have recently been shown to be “an essential non-redundant regulatory subset that supplements nTreg cells, in part by expanding TCR diversity within regulatory responses”.[26] Acute depletion of the iTreg cell pool in mouse models has resulted in inflammation and weight loss. The contribution of nTreg cells versus iTreg cells in maintaining tolerance is unknown, but both are important. Epigenetic differences have been observed between nTreg and iTreg cells, with the former having more stable Foxp3 expression and wider demethylation.
The small intestinal environment is high in vitamin A and is a location where retinoic acid is produced.[27] The retinoic acid and TGF-beta produced by dendritic cells within this area signal for production of regulatory T cells.[27] Vitamin A and TGF-beta promote T cell differentiation into regulatory T cells opposed to Th17 cells, even in the presence of IL-6.[28][29] The intestinal environment can lead to induced regulatory T cells (iTregs) with TGF-beta and retinoic acid[30], some of which express the lectin-like receptor CD161 and are specialized to maintain barrier integrity by accelerating wound healing [31]. The Tregs within the gut are differentiated from naïve T cells after antigen is introduced.[32]
Immune-suppression
An important question in the field of immunology is how the immunosuppressive activity of regulatory T cells is modulated during the course of an ongoing immune response. While the immunosuppressive function of regulatory T cells prevents the development of autoimmune disease, it is not desirable during immune responses to infectious microorganisms. Current hypotheses suggest that, upon encounter with infectious microorganisms, the activity of regulatory T cells may be downregulated, either directly or indirectly, by other cells to facilitate elimination of the infection. Experimental evidence from mouse models suggests that some pathogens may have evolved to manipulate regulatory T cells to immunosuppress the host and so potentiate their own survival. For example, regulatory T cell activity has been reported to increase in several infectious contexts, such as retroviral infections (the most well-known of which is HIV), mycobacterial infections (like tuberculosis), and various parasitic infections including Leishmania and malaria.
Tregs play major roles during HIV infection. They suppress the immune system, thus limiting target cells and reducing inflammation, but this simultaneously disrupts the clearance of virus by the cell-mediated immune response and enhances the reservoir by pushing CD4+ T cells to a resting state, including infected cells. Additionally, Tregs can be infected by HIV, increasing the size of the HIV reservoir directly. Thus, Tregs are being investigated as targets for HIV cure research.[33]
Regulatory T cells have a large role in the pathology of visceral leishmaniasis and in preventing excess inflammation in patients cured of visceral leishmaniasis.
CD4+ regulatory T cells are often associated with solid tumours in both humans and murine models. Increased numbers of regulatory T cells in breast, colorectal and ovarian cancers is associated with a poorer prognosis.[34]
CD70+ non-Hodgkin lymphoma B cells induce Foxp3 expression and regulatory function in intratumoral CD4+CD25− T cells.[35]
A recent study shows that cerebral ischemia can increase bone marrow CD4(+)CD25(+)Foxp3(+) regulatory T cells via signals from the sympathetic nervous system.[36]
Cancer and the Tumor Response
Most tumors elicit an immune response in the host that is mediated by tumor antigens, thus distinguishing the tumor from other non-cancerous cells. This causes large numbers of tumor-infiltrating lymphocytes (TILs) to be found in the tumor microenvironment.[37] Although it is not entirely understood, it is thought that these lymphocytes target cancerous cells and therefore slow or terminate the development of the tumor. However, this process is complicated because Tregs seem to be preferentially trafficked to the tumor microenvironment. While Tregs normally make up only about 4% of CD4+ T cells, they can make up as much as 20–30% of the total CD4+population around the tumor microenvironment.[38]
Although high levels of TILs were initially thought to be important in determining an immune response against cancer, it is now widely recognized that the ratio of Tregs to Teffectors in the tumor microenvironment is a determining factor in the success of the immune response against the cancer. High levels of Tregs in the tumor microenvironment are associated with[7] poor prognosis in many cancers, such as ovarian, breast, renal, and pancreatic cancer.[38] This indicates that Tregs suppress Teffector cells and hinder the body’s immune response against the cancer.
However, because in Science, there are many howevers, in some types of cancer the opposite is true, and high levels of Tregs are associated with a positive prognosis. This trend is seen in cancers such as colorectal carcinoma and follicular lymphoma. This could be due to Treg’s ability to suppress general inflammation which is known to trigger cell proliferation and metastasis .[38] These opposite effects indicate that Treg’s role in the development of cancer is highly dependent on both type and location of the tumor.
Although it is still not entirely understood how Tregs are preferentially trafficked to the tumor microenvironment, the chemotaxis is probably driven by the production of chemokines by the tumor. Treg infiltration into the tumor microenvironment is facilitated by the binding of the chemokine receptor CCR4, which is expressed on Tregs, to its ligand CCL22, which is secreted by many types of tumor cells.[39] Treg expansion at the site of the tumor could also explain the increased levels of Tregs. The cytokine, TGF-β, which is commonly produced by tumor cells, is known to induce the differentiation and expansion of Tregs.[39]
In general, the immunosuppression of the tumor microenvironment has largely contributed to the unsuccessful outcomes of many conventional cancer immunotherapy treatments. Depletion of Tregs in animal models has shown an increased efficacy of immunotherapy treatments, and therefore, many immunotherapy treatments are now incorporating Treg depletion.[2] In holistic oncology, the approach is different (See Master Class)
Thymus
Similar to other T cells, regulatory T cells develop in the thymus. The latest research suggests that regulatory T cells are defined by expression of the forkhead family transcription factor Foxp3 (forkhead box p3). Expression of Foxp3 is required for regulatory T cell development and appears to control a genetic program specifying this cell’s fate.[40] The large majority of Foxp3-expressing regulatory T cells are found within the major histocompatibility complex (MHC) class II restricted CD4-expressing (CD4+) population and express high levels of the interleukin-2 receptor alpha chain (CD25). In addition to the Foxp3-expressing CD4+ CD25+, there also appears to be a minor population of MHC class I restricted CD8+ Foxp3-expressing regulatory T cells. These Foxp3-expressing CD8+ T cells do not appear to be functional in healthy individuals but are induced in autoimmune disease states by T cell receptor stimulation to suppress IL-17-mediated immune responses.[41] Unlike conventional T cells, regulatory T cells do not produce IL-2 and are therefore anergic at baseline.
A number of different methods are employed in research to identify and monitor Treg cells. Originally, high expression of CD25 and CD4 surface markers was used (CD4+CD25+ cells). This is problematic as CD25 is also expressed on non-regulatory T cells in the setting of immune activation such as during an immune response to a pathogen. As defined by CD4 and CD25 expression, regulatory T cells comprise about 5–10% of the mature CD4+ T cell subpopulation in mice and humans, while about 1–2% of Treg can be measured in whole blood. The additional measurement of cellular expression of Foxp3 protein allowed a more specific analysis of Treg cells (CD4+CD25+Foxp3+ cells). However, Foxp3 is also transiently expressed in activated human effector T cells, thus complicating a correct Treg analysis using CD4, CD25 and Foxp3 as markers in humans. Therefore, some research groups use another marker, the absence or low-level expression of the surface protein CD127 in combination with the presence of CD4 and CD25. Several additional markers have been described, e.g., high levels of CTLA-4 (cytotoxic T-lymphocyte associated molecule-4) and GITR (glucocorticoid-induced TNF receptor) are also expressed on regulatory T cells, however the functional significance of this expression remains to be defined. There is a great interest in identifying cell surface markers that are uniquely and specifically expressed on all Foxp3-expressing regulatory T cells. However, to date no such molecule has been identified.
In addition to the search for novel protein markers, a different method to analyze and monitor Treg cells more accurately has been described in the literature. This method is based on DNA methylation analysis. Only in Treg cells, but not in any other cell type, including activated effector T cells, a certain region within the Foxp3 gene (TSDR, Treg-specific-demethylated region) is found demethylated, which allows to monitor Treg cells through a PCR reaction or other DNA-based analysis methods.[42] Interplay between the Th17 cells and regulatory T cells are important in many diseases like respiratory diseases.[43]
Recent evidence suggests that mast cells may be important mediators of Treg-dependent peripheral tolerance.[44]
Regulatory T cell epitopes
Regulatory T cell epitopes (‘Tregitopes’) were discovered in 2008 and consist of linear sequences of amino acids contained within monoclonal antibodies and immunoglobulin G (IgG). Since their discovery, evidence has indicated Tregitopes may be crucial to the activation of natural regulatory T cells.[45][46][46][47]
Potential applications of regulatory T cell epitopes have been hypothesised: tolerisation to transplants, protein drugs, blood transfer therapies, and type I diabetes as well as reduction of immune response for the treatment of allergies.[48][49][50][51][52][53][47]
Genetic Mutations
Genetic mutations in the gene encoding Foxp3 have been identified in both humans and mice based on the heritable disease caused by these mutations. This disease provides the most striking evidence that regulatory T cells play a critical role in maintaining normal immune system function. Humans with mutations in Foxp3 suffer from a severe and rapidly fatal autoimmune disorder known as Immune dysregulation, Polyendocrinopathy, Enteropathy X-linked (IPEX) syndrome.[54][55]
The IPEX syndrome is characterized by the development of overwhelming systemic autoimmunity in the first year of life, resulting in the commonly observed triad of watery diarrhea, eczematous dermatitis, and endocrinopathy seen most commonly as insulin-dependent diabetes mellitus. Most individuals have other autoimmune phenomena including Coombs-positive hemolytic anemia, autoimmune thrombocytopenia, autoimmune neutropenia, and tubular nephropathy. The majority of affected males die within the first year of life of either metabolic derangements or sepsis. An analogous disease is also observed in a spontaneous Foxp3-mutant mouse known as “scurfy”.
Text under investigation and also under construction
- ^ Bettelli E, Carrier Y, Gao W, Korn T, Strom TB, Oukka M, Weiner HL, Kuchroo VK (May 2006). “Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells”. Nature. 441 (7090): 235–8. doi:10.1038/nature04753. PMID 16648838.
- ^ Jump up to: a b Curiel TJ (May 2007). “Tregs and rethinking cancer immunotherapy”. The Journal of Clinical Investigation. 117 (5): 1167–74. doi:10.1172/JCI31202. PMC 1857250. PMID 17476346.
- ^ Chen W (August 2011). “Tregs in immunotherapy: opportunities and challenges”. Immunotherapy. 3 (8): 911–4. doi:10.2217/imt.11.79. PMID 21843075.
- ^ Miyara M, Gorochov G, Ehrenstein M, Musset L, Sakaguchi S, Amoura Z (October 2011). “Human FoxP3+ regulatory T cells in systemic autoimmune diseases”. Autoimmunity Reviews. 10 (12): 744–55. doi:10.1016/j.autrev.2011.05.004. PMID 21621000.
- ^ Nosbaum A, Prevel N, Truong HA, Mehta P, Ettinger M, Scharschmidt TC, Ali NH, Pauli ML, Abbas AK, Rosenblum MD (Mar 2016). “Cutting Edge: Regulatory T Cells Facilitate Cutaneous Wound Healing”. J Immunol. 196 (5): 2010–4. doi:10.4049/jimmunol.1502139. PMC 4761457. PMID 26826250.
- ^ Adeegbe DO, Nishikawa H (2013). “Natural and induced T regulatory cells in cancer”. Frontiers in Immunology. 4: 190. doi:10.3389/fimmu.2013.00190. PMC 3708155. PMID 23874336.
- ^ Jump up to: a b Curiel TJ (April 2008). “Regulatory T cells and treatment of cancer”. Current Opinion in Immunology. 20 (2): 241–6. doi:10.1016/j.coi.2008.04.008. PMC 3319305. PMID 18508251.
- ^ Hori S, Nomura T, Sakaguchi S (February 2003). “Control of regulatory T cell development by the transcription factor Foxp3″. Science. 299 (5609): 1057–61. doi:10.1126/science.1079490. PMID 12522256.
- ^ Singh B, Schwartz JA, Sandrock C, Bellemore SM, Nikoopour E (November 2013). “Modulation of autoimmune diseases by interleukin (IL)-17 producing regulatory T helper (Th17) cells”. The Indian Journal of Medical Research. 138(5): 591–4. PMC 3928692. PMID 24434314.
- ^ Shevach EM (2000). “Regulatory T cells in autoimmmunity*”. Annual Review of Immunology. 18: 423–49. doi:10.1146/annurev.immunol.18.1.423. PMID 10837065.
- ^ Schmetterer, Klaus G.; Neunkirchner, Alina; Pickl, Winfried F. (June 2012). “Naturally occurring regulatory T cells: markers, mechanisms, and manipulation”. The FASEB Journal. 26 (6): 2253–2276. doi:10.1096/fj.11-193672. ISSN 0892-6638. PMID 22362896.
- ^ Sakaguchi S (2004). “Naturally arising CD4+ regulatory t cells for immunologic self-tolerance and negative control of immune responses”. Annual Review of Immunology. 22: 531–62. doi:10.1146/annurev.immunol.21.120601.141122. PMID 15032588.
- ^ Read S, Malmström V, Powrie F (July 2000). “Cytotoxic T lymphocyte-associated antigen 4 plays an essential role in the function of CD25(+)CD4(+) regulatory cells that control intestinal inflammation”. The Journal of Experimental Medicine. 192(2): 295–302. doi:10.1084/jem.192.2.295. PMC 2193261. PMID 10899916.
- ^ Collison LW, Workman CJ, Kuo TT, Boyd K, Wang Y, Vignali KM, Cross R, Sehy D, Blumberg RS, Vignali DA (November 2007). “The inhibitory cytokine IL-35 contributes to regulatory T-cell function”. Nature. 450 (7169): 566–9. doi:10.1038/nature06306. PMID 18033300.
- ^ Annacker O, Asseman C, Read S, Powrie F (June 2003). “Interleukin-10 in the regulation of T cell-induced colitis”. Journal of Autoimmunity. 20 (4): 277–9. doi:10.1016/s0896-8411(03)00045-3. PMID 12791312.
- ^ Kearley J, Barker JE, Robinson DS, Lloyd CM (December 2005). “Resolution of airway inflammation and hyperreactivity after in vivo transfer of CD4+CD25+ regulatory T cells is interleukin 10 dependent”. The Journal of Experimental Medicine. 202 (11): 1539–47. doi:10.1084/jem.20051166. PMC 1350743. PMID 16314435.
- ^ Gondek DC, Lu LF, Quezada SA, Sakaguchi S, Noelle RJ (February 2005). “Cutting edge: contact-mediated suppression by CD4+CD25+ regulatory cells involves a granzyme B-dependent, perforin-independent mechanism”. Journal of Immunology. 174 (4): 1783–6. doi:10.4049/jimmunol.174.4.1783. PMID 15699103.
- ^ Puccetti P, Grohmann U (October 2007). “IDO and regulatory T cells: a role for reverse signalling and non-canonical NF-kappaB activation”. Nature Reviews. Immunology. 7 (10): 817–23. doi:10.1038/nri2163. PMID 17767193.
- ^ Borsellino G, Kleinewietfeld M, Di Mitri D, Sternjak A, Diamantini A, Giometto R, Höpner S, Centonze D, Bernardi G, Dell’Acqua ML, Rossini PM, Battistini L, Rötzschke O, Falk K (August 2007). “Expression of ectonucleotidase CD39 by Foxp3+ Treg cells: hydrolysis of extracellular ATP and immune suppression”. Blood. 110 (4): 1225–32. doi:10.1182/blood-2006-12-064527. PMID 17449799.
- ^ Kobie JJ, Shah PR, Yang L, Rebhahn JA, Fowell DJ, Mosmann TR (November 2006). “T regulatory and primed uncommitted CD4 T cells express CD73, which suppresses effector CD4 T cells by converting 5′-adenosine monophosphate to adenosine”. Journal of Immunology. 177 (10): 6780–6. doi:10.4049/jimmunol.177.10.6780. PMID 17082591.
- ^ Huang CT, Workman CJ, Flies D, Pan X, Marson AL, Zhou G, Hipkiss EL, Ravi S, Kowalski J, Levitsky HI, Powell JD, Pardoll DM, Drake CG, Vignali DA (October 2004). “Role of LAG-3 in regulatory T cells”. Immunity. 21 (4): 503–13. doi:10.1016/j.immuni.2004.08.010. PMID 15485628.
- ^ Yu X, Harden K, Gonzalez LC, Francesco M, Chiang E, Irving B, Tom I, Ivelja S, Refino CJ, Clark H, Eaton D, Grogan JL (January 2009). “The surface protein TIGIT suppresses T cell activation by promoting the generation of mature immunoregulatory dendritic cells”. Nature Immunology. 10 (1): 48–57. doi:10.1038/ni.1674. PMID 19011627.
- ^ Sakaguchi S, Yamaguchi T, Nomura T, Ono M (May 2008). “Regulatory T cells and immune tolerance”. Cell. 133 (5): 775–87. doi:10.1016/j.cell.2008.05.009. PMID 18510923.
- ^ Pandiyan P, Zheng L, Ishihara S, Reed J, Lenardo MJ (December 2007). “CD4+CD25+Foxp3+ regulatory T cells induce cytokine deprivation-mediated apoptosis of effector CD4+ T cells”. Nature Immunology. 8 (12): 1353–62. doi:10.1038/ni1536. PMID 17982458.
- ^ Walker LS, Sansom DM (November 2011). “The emerging role of CTLA4 as a cell-extrinsic regulator of T cell responses”. Nature Reviews. Immunology. 11 (12): 852–63. doi:10.1038/nri3108. PMID 22116087.
- ^ Haribhai D, Williams JB, Jia S, Nickerson D, Schmitt EG, Edwards B, Ziegelbauer J, Yassai M, Li SH, Relland LM, Wise PM, Chen A, Zheng YQ, Simpson PM, Gorski J, Salzman NH, Hessner MJ, Chatila TA, Williams CB (July 2011). “A requisite role for induced regulatory T cells in tolerance based on expanding antigen receptor diversity”. Immunity. 35 (1): 109–22. doi:10.1016/j.immuni.2011.03.029. PMC 3295638. PMID 21723159.
- ^ Jump up to: a b Sun CM, Hall JA, Blank RB, Bouladoux N, Oukka M, Mora JR, Belkaid Y (August 2007). “Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid”. The Journal of Experimental Medicine. 204 (8): 1775–85. doi:10.1084/jem.20070602. PMC 2118682. PMID 17620362.
- ^ Mucida D, Park Y, Kim G, Turovskaya O, Scott I, Kronenberg M, Cheroutre H (July 2007). “Reciprocal TH17 and regulatory T cell differentiation mediated by retinoic acid”. Science. 317 (5835): 256–60. doi:10.1126/science.1145697. PMID 17569825.
- ^ Erkelens MN, Mebius RE (March 2017). “Retinoic Acid and Immune Homeostasis: A Balancing Act”. Trends in Immunology. 38 (3): 168–180. doi:10.1016/j.it.2016.12.006. PMID 28094101.
- ^ Ziegler SF, Buckner JH (April 2009). “FOXP3 and the regulation of Treg/Th17 differentiation”. Microbes and Infection. 11 (5): 594–8. doi:10.1016/j.micinf.2009.04.002. PMC 2728495. PMID 19371792.
- ^ Povoleri, Giovanni A. M.; Nova-Lamperti, Estefania; Scottà, Cristiano; Fanelli, Giorgia; Chen, Yun-Ching; Becker, Pablo D.; Boardman, Dominic; Costantini, Benedetta; Romano, Marco (2018-11-05). “Human retinoic acid–regulated CD161+ regulatory T cells support wound repair in intestinal mucosa”. Nature Immunology. 19 (12): 1403–1414. doi:10.1038/s41590-018-0230-z. ISSN 1529-2908. PMID 30397350.
- ^ Coombes JL, Siddiqui KR, Arancibia-Cárcamo CV, Hall J, Sun CM, Belkaid Y, Powrie F (August 2007). “A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism”. The Journal of Experimental Medicine. 204 (8): 1757–64. doi:10.1084/jem.20070590. PMC 2118683. PMID 17620361.
- ^ Kleinman AJ, Sivanandham R, Pandrea I, Chougnet CA, Apetrei C (2018). “Regulatory T Cells As Potential Targets for HIV Cure Research”. Frontiers in Immunology. 9: 734. doi:10.3389/fimmu.2018.00734. PMC 5908895. PMID 29706961.
- ^ Dranoff G (December 2005). “The therapeutic implications of intratumoral regulatory T cells”. Clinical Cancer Research. 11 (23): 8226–9. doi:10.1158/1078-0432.CCR-05-2035. PMID 16322278.
- ^ Yang ZZ, Novak AJ, Ziesmer SC, Witzig TE, Ansell SM (October 2007). “CD70+ non-Hodgkin lymphoma B cells induce Foxp3 expression and regulatory function in intratumoral CD4+CD25 T cells”. Blood. 110 (7): 2537–44. doi:10.1182/blood-2007-03-082578. PMC 1988926. PMID 17615291.
- ^ Wang J, Yu L, Jiang C, Fu X, Liu X, Wang M, Ou C, Cui X, Zhou C, Wang J (January 2015). “Cerebral ischemia increases bone marrow CD4+CD25+FoxP3+ regulatory T cells in mice via signals from sympathetic nervous system”. Brain, Behavior, and Immunity. 43: 172–83. doi:10.1016/j.bbi.2014.07.022. PMC 4258426. PMID 25110149.
- ^ Gooden MJ, de Bock GH, Leffers N, Daemen T, Nijman HW (June 2011). “The prognostic influence of tumour-infiltrating lymphocytes in cancer: a systematic review with meta-analysis”. British Journal of Cancer. 105 (1): 93–103. doi:10.1038/bjc.2011.189. PMC 3137407. PMID 21629244.
- ^ Jump up to: a b c Oleinika K, Nibbs RJ, Graham GJ, Fraser AR (January 2013). “Suppression, subversion and escape: the role of regulatory T cells in cancer progression”. Clinical and Experimental Immunology. 171 (1): 36–45. doi:10.1111/j.1365-2249.2012.04657.x. PMC 3530093. PMID 23199321.
- ^ Jump up to: a b Lippitz, Bodo E (2013). “Cytokine patterns in patients with cancer: a systematic review”. The Lancet Oncology. 14 (6): e218–e228. doi:10.1016/s1470-2045(12)70582-x. PMID 23639322.
- ^ Marson A, Kretschmer K, Frampton GM, Jacobsen ES, Polansky JK, MacIsaac KD, Levine SS, Fraenkel E, von Boehmer H, Young RA (February 2007). “Foxp3 occupancy and regulation of key target genes during T-cell stimulation”. Nature. 445 (7130): 931–5. doi:10.1038/nature05478. PMC 3008159. PMID 17237765.
- ^ Ellis SD, McGovern JL, van Maurik A, Howe D, Ehrenstein MR, Notley CA. “Induced CD8+FoxP3+ Treg cells in rheumatoid arthritis are modulated by p38 phosphorylation and monocytes expressing membrane tumor necrosis factor α and CD86.”. 2014. Arthritis Rheumatol. 66(10):2694-705. doi:10.1002/art.38761. PMID 24980778
- ^ Wieczorek G, Asemissen A, Model F, Turbachova I, Floess S, Liebenberg V, Baron U, Stauch D, Kotsch K, Pratschke J, Hamann A, Loddenkemper C, Stein H, Volk HD, Hoffmüller U, Grützkau A, Mustea A, Huehn J, Scheibenbogen C, Olek S (January 2009). “Quantitative DNA methylation analysis of FOXP3 as a new method for counting regulatory T cells in peripheral blood and solid tissue”. Cancer Research. 69 (2): 599–608. doi:10.1158/0008-5472.CAN-08-2361. PMID 19147574.
- ^ Agarwal A, Singh M, Chatterjee BP, Chauhan A, Chakraborti A (2014). “Interplay of T Helper 17 Cells with CD4(+)CD25(high) FOXP3(+) Tregs in Regulation of Allergic Asthma in Pediatric Patients”. International Journal of Pediatrics. 2014: 1–8. doi:10.1155/2014/636238. PMC 4065696. PMID 24995020.
- ^ Lu LF, Lind EF, Gondek DC, Bennett KA, Gleeson MW, Pino-Lagos K, Scott ZA, Coyle AJ, Reed JL, Van Snick J, Strom TB, Zheng XX, Noelle RJ (August 2006). “Mast cells are essential intermediaries in regulatory T-cell tolerance”. Nature. 442(7106): 997–1002. doi:10.1038/nature05010. PMID 16921386.
- ^ Tregitopes can dampen unwanted immune responses through selective activation. “Tregitopes”. EpiVax, Inc.
- ^ Jump up to: a b Hui DJ, Basner-Tschakarjan E, Chen Y, Davidson RJ, Buchlis G, Yazicioglu M, Pien GC, Finn JD, Haurigot V, Tai A, Scott DW, Cousens LP, Zhou S, De Groot AS, Mingozzi F (September 2013). “Modulation of CD8+ T cell responses to AAV vectors with IgG-derived MHC class II epitopes”. Molecular Therapy. 21 (9): 1727–37. doi:10.1038/mt.2013.166. PMC 3776637. PMID 23857231.
- ^ Jump up to: a b De Groot AS, Moise L, McMurry JA, Wambre E, Van Overtvelt L, Moingeon P, Scott DW, Martin W (October 2008). “Activation of natural regulatory T cells by IgG Fc-derived peptide “Tregitopes““. Blood. 112 (8): 3303–11. doi:10.1182/blood-2008-02-138073. PMC 2569179. PMID 18660382.
- ^ “New $2.25M infusion of NIH funds for EpiVax’ Tregitope, proposed “Paradigm-Shifting” Treatment”. Fierce Biotech Research.
- ^ Su Y, Rossi R, De Groot AS, Scott DW (August 2013). “Regulatory T cell epitopes (Tregitopes) in IgG induce tolerance in vivo and lack immunogenicity per se”. Journal of Leukocyte Biology. 94 (2): 377–83. doi:10.1189/jlb.0912441. PMC 3714563. PMID 23729499.
- ^ Cousens LP, Su Y, McClaine E, Li X, Terry F, Smith R, Lee J, Martin W, Scott DW, De Groot AS (2013). “Application of IgG-derived natural Treg epitopes (IgG Tregitopes) to antigen-specific tolerance induction in a murine model of type 1 diabetes”. Journal of Diabetes Research. 2013: 1–17. doi:10.1155/2013/621693. PMC 3655598. PMID 23710469.
- ^ Cousens LP, Mingozzi F, van der Marel S, Su Y, Garman R, Ferreira V, Martin W, Scott DW, De Groot AS (October 2012). “Teaching tolerance: New approaches to enzyme replacement therapy for Pompe disease”. Human Vaccines & Immunotherapeutics. 8 (10): 1459–64. doi:10.4161/hv.21405. PMC 3660767. PMID 23095864.
- ^ Cousens LP, Najafian N, Mingozzi F, Elyaman W, Mazer B, Moise L, Messitt TJ, Su Y, Sayegh M, High K, Khoury SJ, Scott DW, De Groot AS (January 2013). “In vitro and in vivo studies of IgG-derived Treg epitopes (Tregitopes): a promising new tool for tolerance induction and treatment of autoimmunity”. Journal of Clinical Immunology. 33 Suppl 1 (1): S43–9. doi:10.1007/s10875-012-9762-4. PMC 3538121. PMID 22941509.
- ^ Elyaman W, Khoury SJ, Scott DW, De Groot AS (2011). “Potential application of tregitopes as immunomodulating agents in multiple sclerosis”. Neurology Research International. 2011: 1–6. doi:10.1155/2011/256460. PMC 3175387. PMID 21941651.
- ^ Online Mendelian Inheritance in Man IPEX
- ^ ipex at NIH/UW GeneTests
- Regulatory+T-Cells at the US National Library of Medicine Medical Subject Headings (MeSH)